
This explanatory animation illustrates SomaLogic’s SOMAmer technology. At the heart of SomaLogic’s unique platform are SOMAmer (Slow Off-rate Modified Aptamer) modified nucleic acid-based protein binding reagents, each of which are highly specific for their cognate protein. To date, SomaLogic has developed SOMAmers to a broad array of over 1000 different protein targets critical to normal and disease biology, and continue to add new SOMAmers all the time.
SOMAscan™ technology (illustrated below) takes advantage of two distinctive properties of SOMAmers: Their specific protein-binding properties and their primary nucleic acid sequences. These two properties not only ensure accurate protein detection and measurement, but allow the multiplexing of literally up to more than a thousand such measurements in each of several hundred different samples in a single experiment.

This technology ensures that measurements are taken only of proteins specific to the SOMAmers being used, giving an accurate readout of both protein identity and concentration in the original sample. The ability to detect anywhere from a few to more than a thousand proteins in literally hundreds of different samples a day – depending on the requirements of the analysis being undertaken – provides for a quick determination of a protein biomarker “signature” indicative of the disease or biological state (e.g., drug response) being studied.
SomaLogic Somamer MOA from Fairman Studios on Vimeo.
Bacteriorhodopsin is a protein used by Archaea, the most notable one being Halobacteria. It acts as a proton pump; that is, it captures light energy and uses it to move protons across the membrane out of the cell.[1] The resulting proton gradient is subsequently converted into chemical energy.[2]
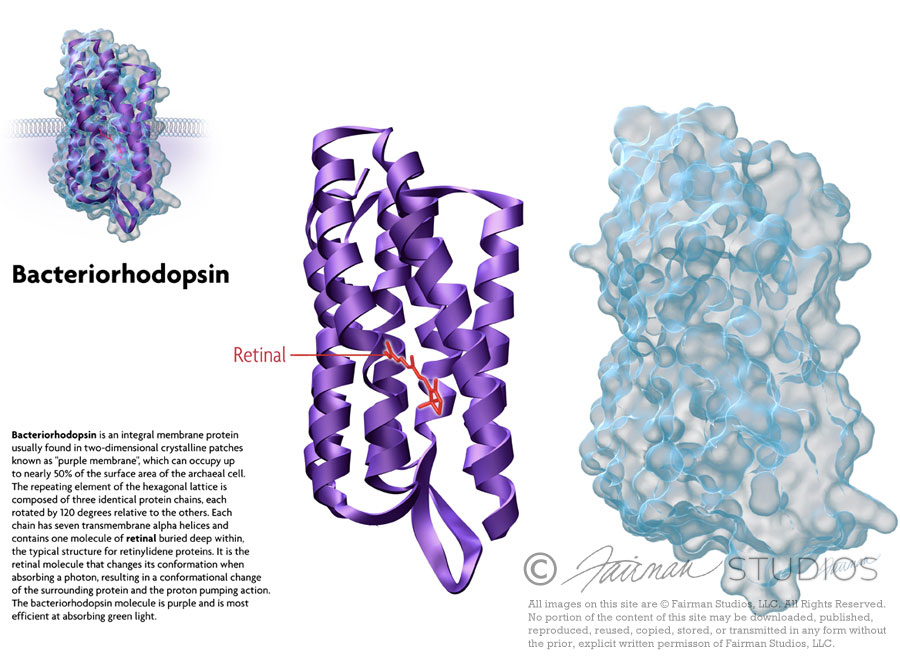
Bacteriorhodopsin is an integral membrane protein usually found in two-dimensional crystalline patches known as “purple membrane”, which can occupy up to nearly 50% of the surface area of the archaeal cell. The repeating element of the hexagonal lattice is composed of three identical protein chains, each rotated by 120 degrees relative to the others. Each chain has seven transmembrane alpha helices and contains one molecule of retinal buried deep within, the typical structure for retinylidene proteins.
It is the retinal molecule that changes its conformation when absorbing a photon, resulting in a conformational change of the surrounding protein and the proton pumping action.[3] It is covalently linked to Lys216 in the chromophore by Schiff base action. After photoisomerization of the retinal molecule, Asp85 becomes a proton acceptor of the donor proton from the retinal molecule. This releases a proton from a “holding site” into the extracellular side (EC) of the membrane. Reprotonation of the retinal molecule by Asp96 restores its original isomerized form. This results in a second proton being released to the EC side. Asp85 releases its proton into the “holding site,” where a new cycle may begin.
The bacteriorhodopsin molecule is purple and is most efficient at absorbing green light (wavelength 500-650 nm, with the absorption maximum at 568 nm).
Bacteriorhodopsin belongs to a family of bacterial proteins related to vertebrate rhodopsins, the pigments that sense light in the retina. Rhodopsins also contain retinal; however, the functions of rhodopsin and bacteriorhodopsin are different, and there is only slight homology in their amino acid sequences. Both rhodopsin and bacteriorhodopsin belong to the 7TM receptor family of proteins, but rhodopsin is a G protein-coupled receptor and bacteriorhodopsin is not. In the first use of electron crystallography to obtain an atomic-level protein structure, the structure of bacteriorhodopsin was resolved in 1990. It was then used as a template to build models of G protein-coupled receptors before crystallographic structures were also available for these proteins.
Many molecules have homology to bacteriorhodopsin, including the light-driven chloride pump halorhodopsin (for which the crystal structure is also known), and some directly light-activated channels like channelrhodopsin.
All other photosynthetic systems in bacteria, algae, and plants use chlorophylls or bacteriochlorophylls rather than bacteriorhodopsin. These also produce a proton gradient, but in a quite different and more indirect way involving an electron transfer chain consisting of several other proteins. Furthermore, chlorophylls are aided in capturing light energy by other pigments known as “antennas”; these are not present in bacteriorhodopsin-based systems. Last, chlorophyll-based photosynthesis is coupled to carbon fixation (the incorporation of carbon dioxide into larger organic molecules); this is not true for bacteriorhodopsin-based system. Thus, it is likely that photosynthesis independently evolved at least twice, once in bacteria and once in archaea.
Sources:



